1. Introduction
2. Materials and methods
2.1 Animals and study setting
2.2 Donor synchronization, superovulation, and artificial insemination
2.3 Recovery of embryos and endometrial epithelial cells
2.4 RNA extraction and cDNA synthesis
2.5 RAPD-PCR fingerprinting and clustering analysis
2.6 Quantitative PCR for embryo-quality–associated transcripts
2.7 ELISA for uterine-fluid hormones
2.8 Statistical analysis
3. Results
3.1 Embryo recovery and grouping of donors/EECs
3.2 Morphological assessment of embryos and EECs
3.3 RAPD-PCR banding profiles and UPGMA-based clustering
3.4 Hormone concentrations in uterine fluid
4. Discussion
5. Conclusion
1. Introduction
Superovulation followed by embryo transfer (ET) is widely used to accelerate genetic dissemination by increasing the number of offspring produced per female within a limited reproductive window. While this approach cannot fully match the genetic output achievable through extensive use of elite sires, it substantially expands the reproductive contribution of valuable donor females beyond what is feasible with a single annual calving cycle [1]. Nevertheless, ET outcomes remain variable. Success is influenced by multiple biological and managerial factors, including embryo quality, seasonality, donor breed and age, nutritional and health status, stress, the superovulation protocol, and procedural proficiency [1].
In mammals, implantation is a sequential process involving blastocyst hatching, intrauterine movement, apposition and attachment to the endometrium, invasion, and subsequent placentation. A considerable proportion of fertilized preimplantation embryos fail to implant [2]. Because implantation occurs within the endometrium, the receptivity of this tissue, shaped by cyclical endocrine cues, is critical for embryo survival and developmental competence [3]. Gonadotropins such as FSH, LH, and chorionic gonadotropin (hCG) are central components of reproductive endocrine regulation [4], and ovarian steroidogenesis is modulated through FSH-driven aromatase activity that converts androgens to estrogens [5]. EECs provide a practical in vitro system to interrogate embryo–uterine signaling pathways, including mechanisms related to interferon-tau–mediated modulation of oxytocin-induced prostaglandin F2α (PGF2α) responses. In addition, the extracellular matrix, composed of collagens, heparan sulfate proteoglycans, laminins, and other components, forms a structural microenvironment that supports embryo development [6]. Beyond classical pregnancy contexts, PAPP-A has been reported in non-pregnant tissues and in males [7], and pregnancy-associated antigens including PAPP-A have also been characterized in primate models using immunoassays based on human-derived antigens [8]. Therefore, this study aimed to evaluate the association between uterine endocrine factors (FSH and LH) and the relative expression of hormone- and implantation-related transcripts in bovine EECs, and to examine how these profiles relate to embryo quality.
2. Materials and methods
2.1 Animals and study setting
All data were obtained from an embryo transfer program performed at a commercial Korean native cattle farm. Donor females were enrolled when they met standard eligibility criteria for ET, including appropriate body condition score and postpartum interval, regular estrous cyclicity, absence of uterine abnormalities, and overall nutritional and health suitability for superovulation and embryo recovery.
2.2 Donor synchronization, superovulation, and artificial insemination
Donors received follicle-stimulating hormone (FSH; Antorin, Kawasaki, Japan) at a total dose of 30 AU, administered every 12 h for four consecutive days beginning 4–5 days after estrus. A progesterone-releasing intravaginal device (CIDR-PLUS; Bioniche Animal Health, Australia) was inserted using a CIDR applicator. PGF2α was administered two days after initiation of the FSH regimen, and the CIDR-PLUS device was removed after four days. Ovulation was induced using GnRH, and frozen–thawed semen was used for AI on three occasions at 12 h intervals (Table 1).
Table 1.
Superovulation, insemination, and uterine flushing schedule used for embryo recovery from donor cows
| Treatment day | Time | Hormonal treatment |
| 0 | 7 AM |
- P4 device insertion - 50 mg P4 - 1 mg E2 |
| 4 | 7 AM | - 6 mg FSH |
| 7 PM | - 6 mg FSH | |
| 5 | 7 AM | - 5 mg FSH |
| 7 PM | - 5 mg FSH | |
| 6 | 7 AM |
- 4 mg FSH - 30 mg PGF2α (Dinoprost) |
| 7 PM |
- 4 mg FSH - 30 mg PGF2α (Dinoprost) | |
| 7 | 7 AM |
- P4 device removal - 3 mg FSH |
| 7 PM | - 3 mg FSH | |
| 8 | 7 AM | - Estrus |
| 7 PM |
- AI - 1 mg GnRH | |
| 9 | 7 AM | - AI |
| 7 PM | - AI | |
| 15 | 7 AM | - Flushing |
2.3 Recovery of embryos and endometrial epithelial cells
Embryos were collected on day 7 after insemination via transcervical uterine flushing with Dulbecco’s phosphate-buffered saline (D-PBS). For the procedure, local anesthesia was provided by injecting 2% lidocaine (Jeil Pharm, Korea) between the first and second lumbar vertebrae, and flushing was conducted using a Foley catheter. Recovered embryos were evaluated using the International Embryo Technology Society (IETS) grading system. EECs were collected from Korean native cattle (n = 10) during the same nonsurgical uterine lavage performed on day 7 after ovulation. Approximately 20 ml of the EEC-containing fraction remaining in the collection cup was transferred to a Petri dish and examined under a stereomicroscope (10×). Red blood cell removal treatment was applied for 10 min, after which EECs were washed twice with EEC holding medium to remove residual debris.
2.4 RNA extraction and cDNA synthesis
Total RNA was extracted from isolated EECs using TRIzol reagent (Invitrogen, USA) following the manufacturer’s protocol. RNA quantity and purity were assessed by absorbance at 260/280 nm using a Beckman DU 600 spectrophotometer (Beckman Instruments, San Ramon, CA). First-strand cDNA was synthesized using SuperScript II reverse transcriptase. Briefly, RNA/primer mixtures were prepared with 8.5 µg total RNA, 1 µL 10 mM dNTP mix, and 0.5 µg oligo(dT), heated at 65°C for 5 min, and immediately chilled on ice for at least 1 min. A separate mixture containing 2 µL 10× RT buffer, 4 µL 5 mM MgCl₂, 2 µL 0.1 M DTT, and 1 µL RNaseOUT was prepared; 9 µL of this mixture was added to each RNA/primer tube and incubated at 42°C for 2 min. SuperScript II RT (0.5 µl; 50 units) was then added, followed by incubation at 42°C for 50 min. Reverse transcription was terminated at 70°C for 15 min, and RNase H (1 µl) was added with an additional incubation at 37°C for 20 min.
2.5 RAPD-PCR fingerprinting and clustering analysis
For RAPD-PCR, 11 UFPF primers (JK, Anseong, KOR) with GC content greater than 60% were used. Each 25 µL PCR reaction contained 1 µL cDNA template, 2.5 µL 10× buffer, 2.5 mM dNTPs, 20 pmol primer, 2 units Taq DNA polymerase, and nuclease-free water to volume. Cycling conditions consisted of an initial denaturation at 95°C for 7 min, 40 cycles of 95°C for 1 min, 58°C for 1 min, and 72°C for 1 min, and a final extension at 72°C for 7 min, followed by a 4°C hold. PCR products were resolved on 1.5% agarose gels at 100 V for 3 h. Banding profiles from good, fair, and outcome groups were used to construct dendrograms. Genetic clustering was performed in MEGA (MEGA5.05 stated in the Methods) using the unweighted pair group method with arithmetic mean (UPGMA), following Tamura et al.
2.6 Quantitative PCR for embryo-quality–associated transcripts
For qPCR analysis, RNA extracted from EECs using TRIzol was treated with DNase (Ambion, Austin, TX) per the manufacturer’s instructions. cDNA was used as a template in SYBR Green master mix (TOYOBO, Japan). Target genes were amplified using gene-specific primers (Table 3), with 18S rRNA as the reference gene. Amplification was performed for 30 cycles, with annealing temperatures ranging from 51 to 65°C, and data were processed using Rotor-Gene Real-Time Software v6.0.
2.7 ELISA for uterine-fluid hormones
For ELISA, protein samples were diluted with 50% assay buffer. Concentrations of FSH, LH, and pregnancy-associated plasma protein-A (PAPP-A) were measured using a sandwich ELISA kit (R&D Systems Europe, Abingdon, UK) according to the manufacturer’s protocol. All assays were run in duplicate and averaged. Hormone concentrations were calculated using a standard curve generated from seven standards and fitted with a four-parameter logistic model, y=(A−D)/[1+(x/C)B]+D. Final values were reported as ng/ml.
2.8 Statistical analysis
ELISA data were analyzed using SAS (version 9.4; SAS Institute Inc., Cary, NC, USA). Differences were considered significant at p < 0.05
3. Results
3.1 Embryo recovery and grouping of donors/EECs
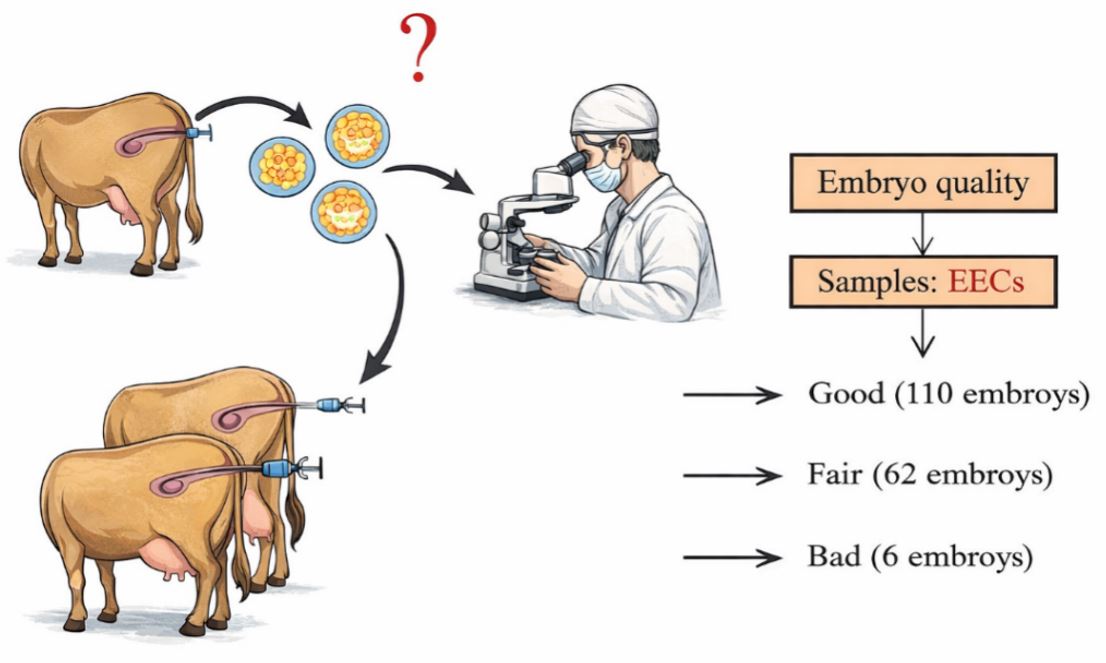
In vivo–derived embryos were produced during April–June 2017 and recovered on day 7 after the first AI. Embryos, EECs, and uterine fluid were obtained by transcervical uterine flushing with D-PBS (Fig. 1). A total of 178 embryos were collected and summarized by developmental stage (morula, compact morula, early blastocyst, and expanded blastocyst) (Table 2). Across the three donor outcome groups defined by embryo quality (good, fair, and bad), the average numbers of embryos recovered were 110, 62, and 6, respectively, and degenerated embryos were observed in each group (Table 2).
Table 2.
Developmental-stage distribution of day 7 embryos recovered from donor cattle (total n = 178), including degenerated embryos, summarized by donor outcome group
|
No. of donors |
No. of embryos | Degenerated | No. of embryos (%) | ||||
| M¹ | CM² | EB³ | EDB⁴ | Subtotal | |||
| 4 | 110 | 22 (20) | 5(5) | 72(65.4) | 3(2.7) | 8(8) | 88 (80) |
| 4 | 62 | 54 (87) | - | 7(11.2) | 1(1.6) | - | 8 (13) |
| 2 | 6 | 4 (66.6) | - | 2 (33.4) | - | - | 2 (33.4) |
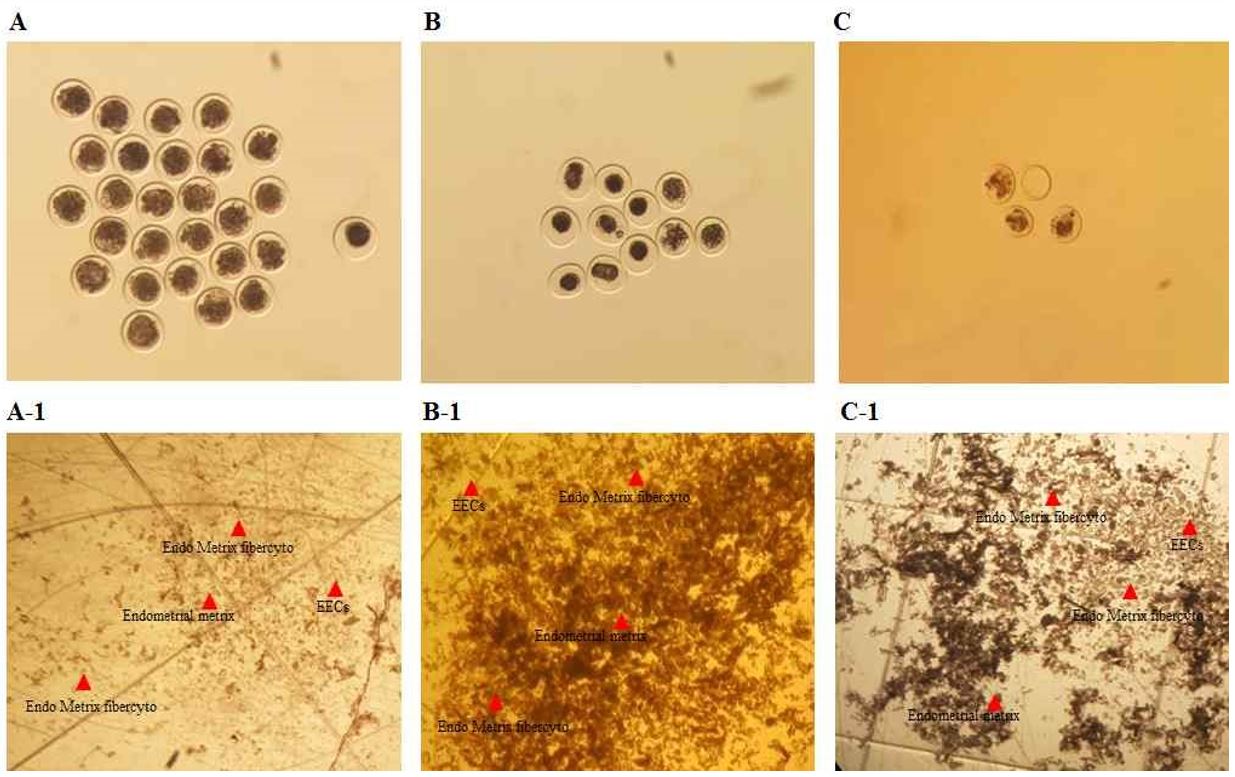
3.2 Morphological assessment of embryos and EECs
Representative morphology of embryos and their corresponding EECs collected after insemination is shown in Fig. 2. Donors were categorized according to embryo quality (good, fair, and bad), and EEC samples recovered from the same uterine lavage were assigned to the corresponding groups (Fig. 2).
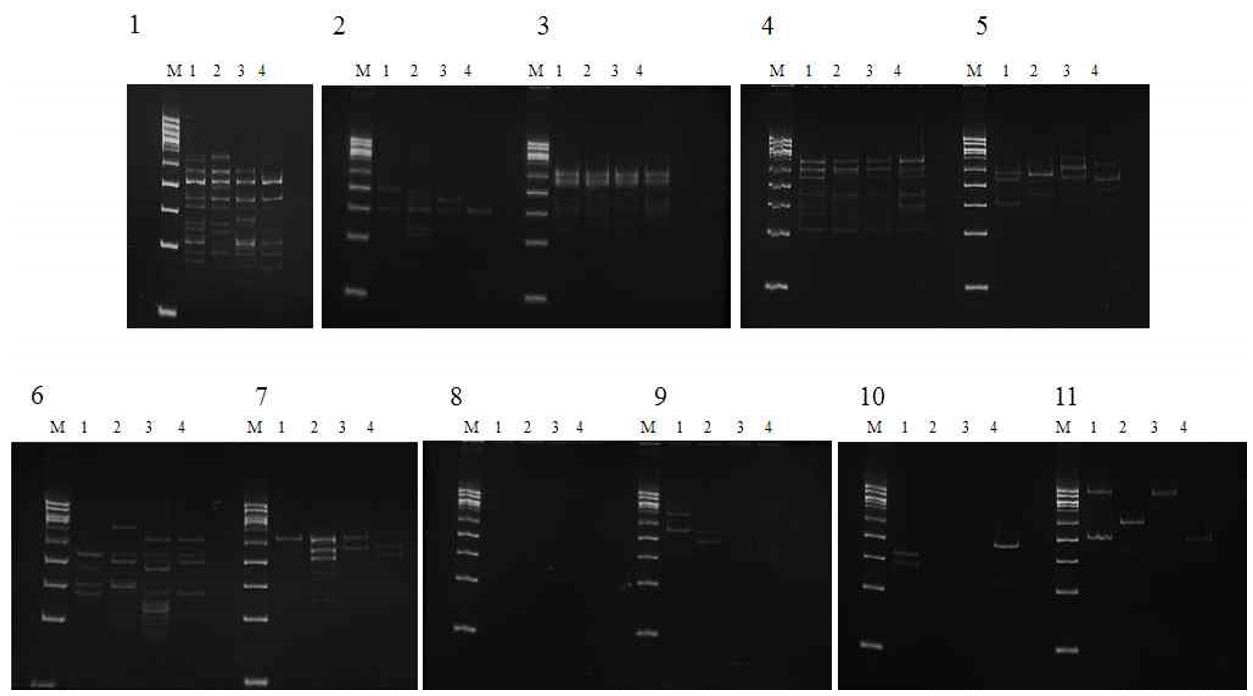
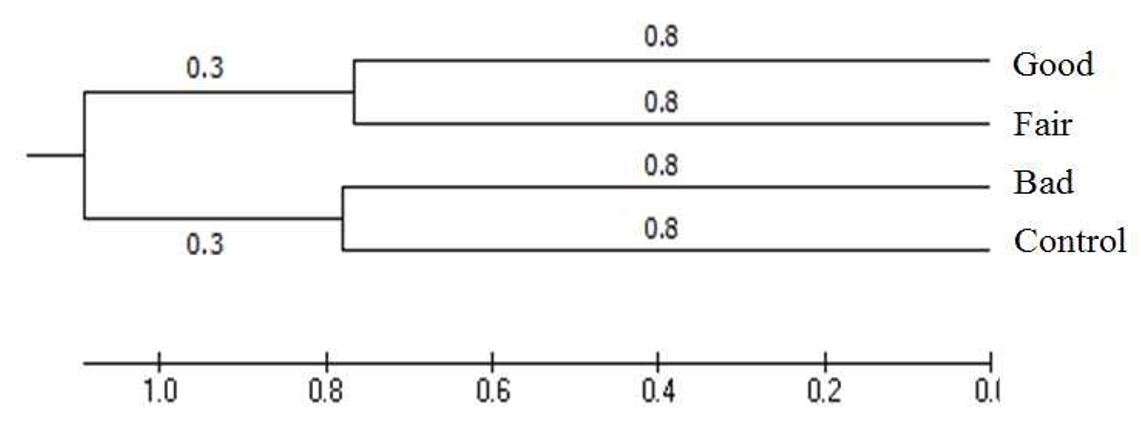
3.3 RAPD-PCR banding profiles and UPGMA-based clustering
RAPD-PCR analysis using cDNA from EECs generated multiple bands spanning approximately 500–1,000 bp across the 11 UFPF primers, with primer-dependent differences in band presence and intensity among the outcome groups (Fig. 3; Tables 3 and 4). When combined RAPD profiles were subjected to UPGMA clustering (Fig. 4), group-level separation was observed. The genetic distance was 0.8 between the good and fair groups and 0.8 between the control and groups (Fig. 4).
Table 3.
RAPD-PCR polymorphism matrix (presence or absence of bands) obtained from endometrial epithelial cell (EEC) cDNA using 11 UFPF primers across donor outcome groups
| Sample | UFPF primers | |||||||||||
| 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9 | 10 | 11 | Total | |
| Good | 1 | - | 4 | - | - | - | - | - | 2 | 2 | - | 9 |
| Fair | 2 | 1 | 3 | 2 | - | - | - | - | 1 | - | 1 | 10 |
| bad | 2 | 3 | - | - | 1 | - | 2 | - | - | - | - | 8 |
| Control | - | - | - | - | - | - | - | - | 1 | 2 | - | 3 |
| Total | 5 | 4 | 7 | 2 | 1 | - | 2 | - | 4 | 4 | 1 | 30 |
Table 4.
Summary of group-discriminatory RAPD-PCR bands detected with UFPF primers in EEC cDNA from good, fair, and bad outcome groups, including approximate amplicon sizes and band counts
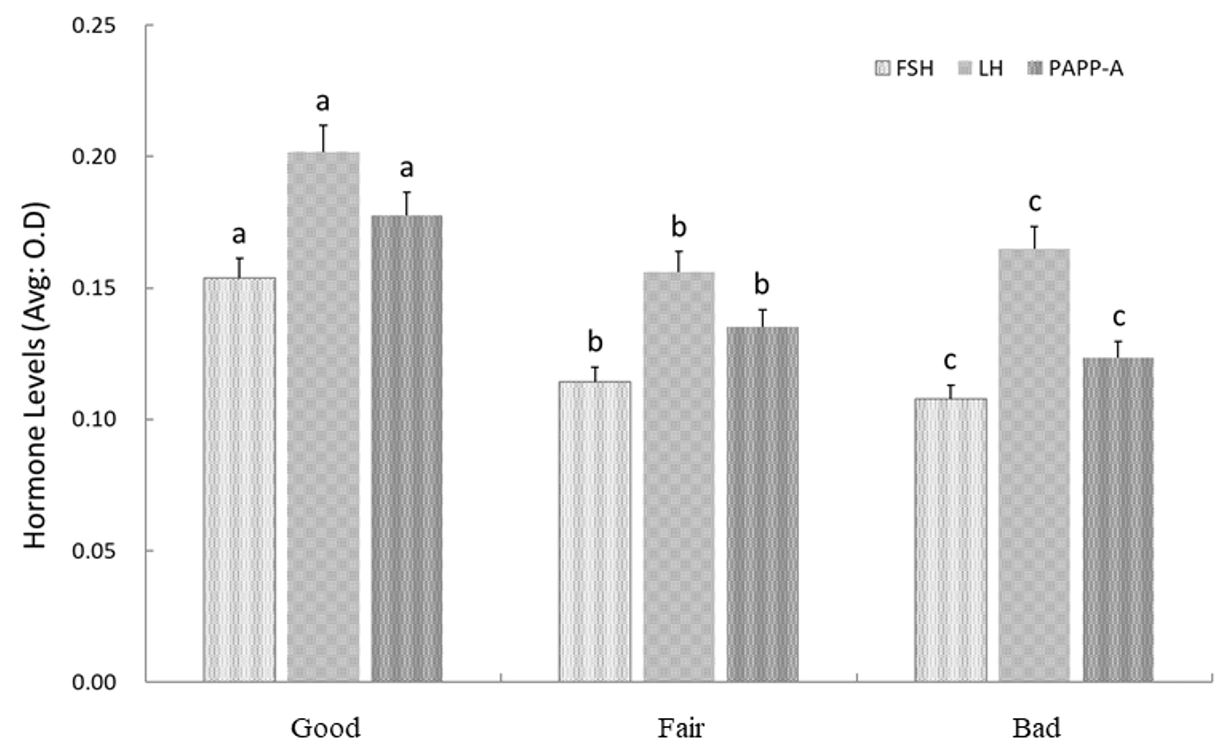
3.4 Hormone concentrations in uterine fluid
FSH, LH, and PAPP-A were detectable in uterine fluid samples from the good, fair, and bad groups (Fig. 5). However, the group exhibited reduced FSH and PAPP-A signals in uterine fluid compared with the other groups. In addition, while the good and fair groups showed broadly comparable abundance patterns for FSH and LH, the group showed lower levels.
4. Discussion
Embryo transfer outcomes are strongly influenced by embryo quality, and morphological assessment remains a practical basis for selection in MOET programs [1]. In the present study, donors were categorized by the day 7 embryo-quality outcome, and EECs collected from the same uterine environment were assigned to the corresponding groups. This paired design enabled comparison of uterine epithelial molecular profiles across distinct embryo-quality phenotypes observed under field ET conditions [1].
Implantation success depends on coordinated communication between the conceptus and the uterine endometrium, and early developmental loss is common in mammals [2]. Therefore, uterine conditions at the time of embryo development can be expected to associate with subsequent embryo quality and developmental competence [2]. In this context, gonadotropin-related endocrine signaling is a central component of reproductive regulation, and fluctuations in such signals can shape the uterine milieu relevant to embryo development [4].
A major observation of this study is that cDNA fingerprinting of EECs produced group-dependent banding patterns that distinguished donor categories defined by embryo-quality outcomes. Although RAPD-PCR is not a transcript-identification method, the consistent separation of banding profiles supports the presence of outcome-associated differences in the uterine epithelial transcript landscape. Notably, the present endocrine profiling further showed reduced FSH and PAPP-A concentrations in uterine fluid from the outcome group. These patterns are consistent with the concept that uterine endocrine status and epithelial molecular features jointly reflect conditions linked to embryo quality [4,6,7].
PAPP-A has been studied primarily in the context of early pregnancy biology and has also been evaluated as part of pregnancy-associated antigen panels in primate models [7,8]. In the present work, lower PAPP-A levels in uterine fluid from the outcome group suggest that variation in this factor, together with gonadotropin-related measures, may be associated with the uterine environment supporting high-quality embryo development [4,7,8]. In addition, clustering analysis was performed using MEGA5 as described previously [9], providing a standardized computational framework for group-level separation of the RAPD-derived profiles.
This study has clear limitations that should be addressed to strengthen translational value. First, RAPD-PCR provides pattern-based discrimination rather than direct identification of the underlying transcripts; therefore, follow-up work should identify the specific sequences contributing to discriminatory bands and validate them using targeted assays in independent cohorts. Second, because the current reference list does not include an IETS guideline citation, a proper reference for embryo grading standards should be added to support statements regarding grading criteria. Finally, integrating validated epithelial markers with uterine fluid endocrine measures will require prospective testing to determine whether combined readouts improve prediction of MOET outcomes beyond either measure alone [1,4].
5. Conclusion
RAPD-PCR–based cDNA fingerprinting of endometrial epithelial cells distinguished donor groups categorized by embryo quality outcome. In addition, uterine fluid from the outcome group showed lower FSH and PAPP-A levels than samples from the good and fair groups. These results indicate that integrating EEC molecular fingerprints with uterine fluid endocrine profiling may provide complementary indicators associated with bovine embryo quality. However, because RAPD-PCR primarily provides pattern-based discrimination, further work is required to identify the underlying transcripts and validate robust marker candidates for practical MOET optimization.